Initial commit
parents
Showing
LICENSE
0 → 100755
README.md
0 → 100755
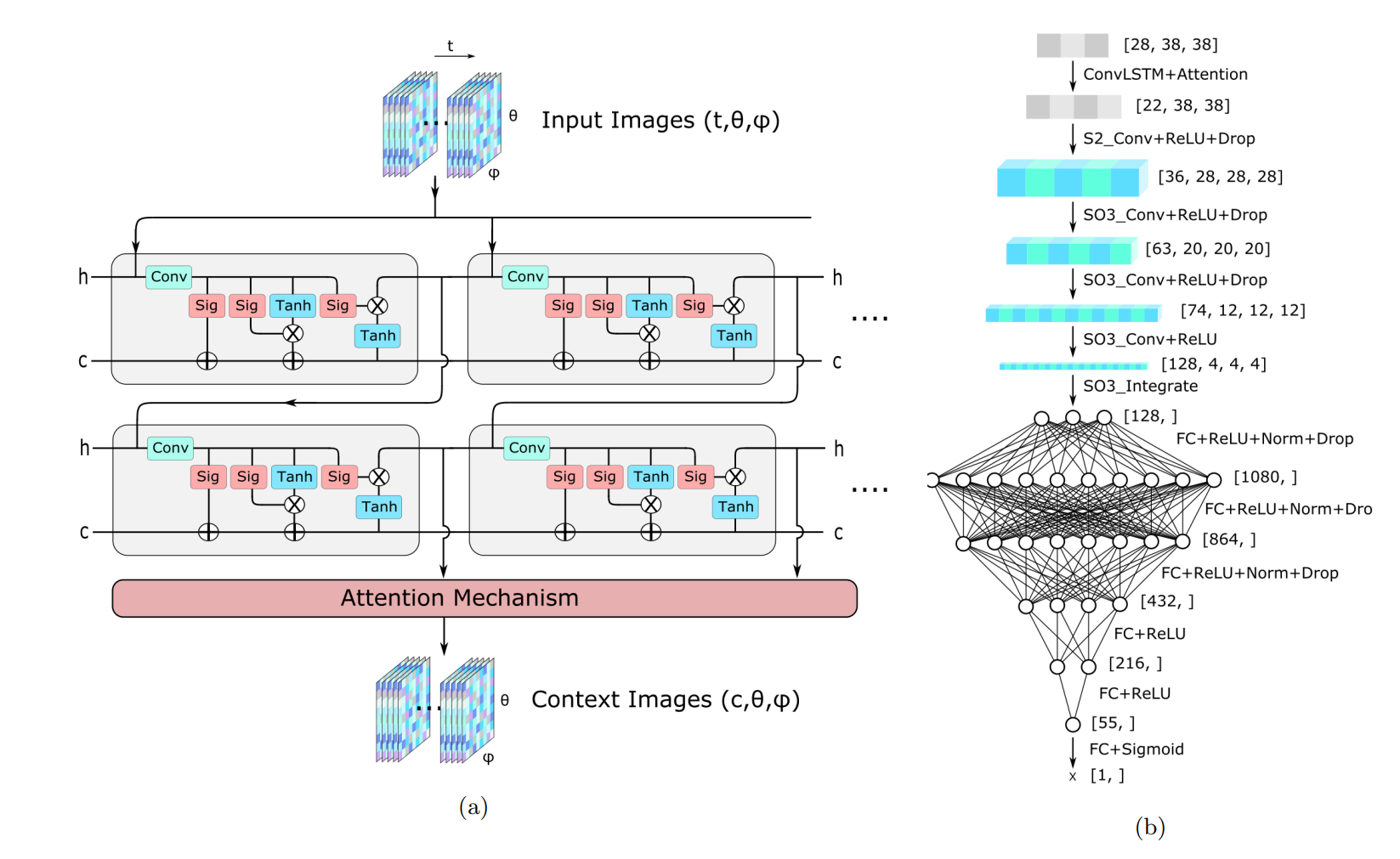
assets/architectures.png
0 → 100755
293 KB
data/clock.py
0 → 100755
data/generate_kamland_mc.py
0 → 100755
data/generate_pickle_list.py
0 → 100755
data/process_kamland.sh
0 → 100755
data/settings.py
0 → 100755
data/tools.py
0 → 100755
docker/.gitkeep
0 → 100755
docker/dockerfile
0 → 100755
lie_learn/.gitignore
0 → 100755
lie_learn/LICENSE
0 → 100755
lie_learn/MANIFEST.in
0 → 100755
lie_learn/MANYLINUX.md
0 → 100755
lie_learn/README.md
0 → 100755