"git@developer.sourcefind.cn:jerrrrry/infinicore.git" did not exist on "d4738a98caafdf96016aaf7f2611a41c428419b4"
Merge pull request #165 from sshaoshuai/master
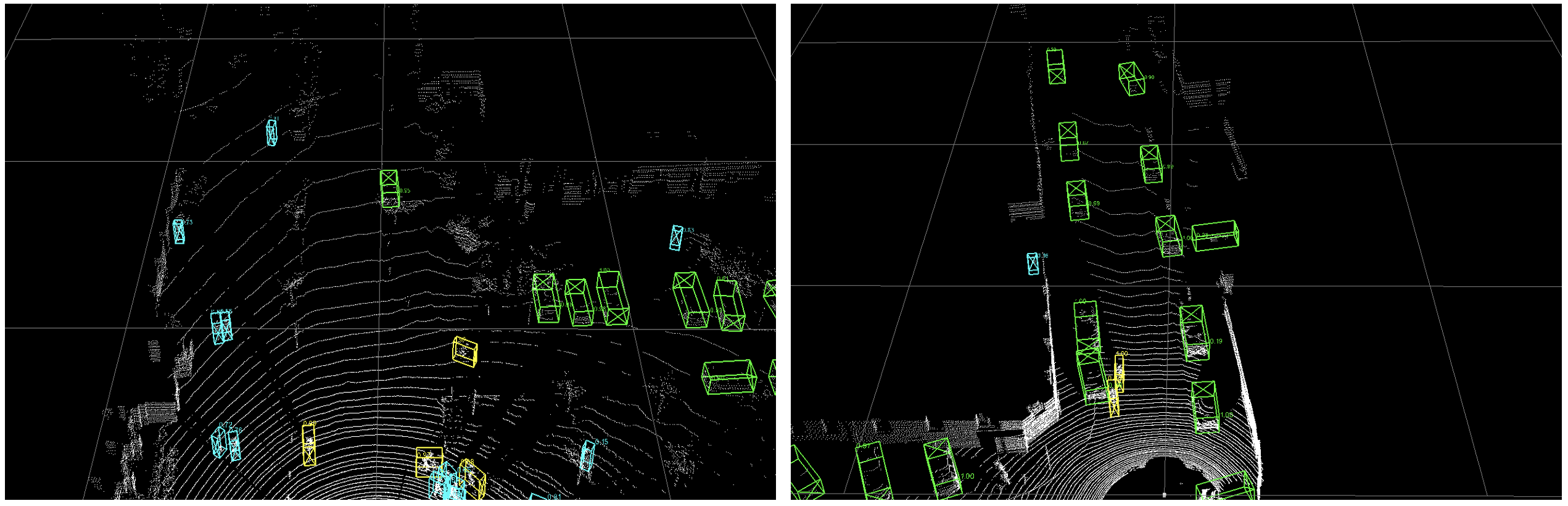
Add visualization codes and a quick demo.
Showing
docs/DEMO.md
0 → 100644
docs/demo.png
0 → 100644
606 KB
tools/demo.py
0 → 100644